Events
Include expired: true
and Keywords: Abiotic stress or DeepMind or EeLP or Intensities or Linux or Medicine or eLearning or lifescience
and Scientific topics: Computational biology or Data management or Literature and language or Workflows
Email Subscription
Subscribe to Filter
Importing into Google Calendar
In the left-hand column of the Google Calendar main view, click the arrow to the right of "Other calendars" and click "Add by URL". In the form that appears, paste in the URL from the box above, and click the button to confirm.
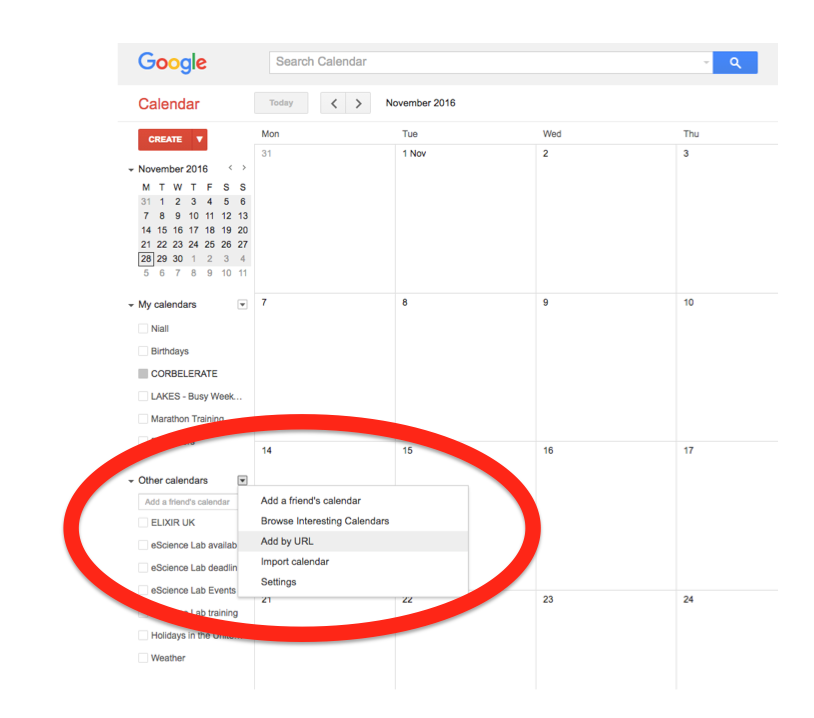
Please note, it may take a while for newly created events in TeSS to synchronise with your Google Calendar.
-
Face-to-faceMeetings and conferencesAustralasian Genomic Technologies Conference
11 - 14 October 2015
Lovedale, Australia
Face-to-face
Computational biology Genomics Bioinformatics ABR lifescience -
Face-to-faceWorkshops and coursesIntroduction to Unix for life scientists
2 June 2016 @ 09:00 - 00:00
Lab-14\, The University of Melbourne, Australia
Face-to-face
Software engineering Computational biology Bioinformatics lifescience ABR VLSCI -
Face-to-faceWorkshops and coursesOpen source science with Git and GitHub
2 August 2016 @ 09:00 - 00:00
University of Melbourne, Australia
Face-to-face
Data submission, annotation, and curation Data management Bioinformatics ABR VLSCI lifescience -
Face-to-faceWorkshops and coursesELIXIR-EXCELERATE HPC Train-the-Researcher course
6 - 7 April 2017
Málaga, Spain
Face-to-face
Computer science Computational biology high-performance computing training eLearning EeLP -
OnlineWorkshops and coursesH3ABioNet 2018 Genotyping Chip Data Analysis and GWAS lecture series - Lecture 3
27 August 2018 @ 15:00 - 16:00
Online
Computational biology Genomics Genotyping experiment Bioinformatics Population genomics Genotype calling H3ABioNet Genome Studio H3Africa genotyping array African populations Array processing Illumina arrays File formats Annotations Non-polymorphic Top/Bottom annotation Genotype array probes Intensities Convert to PLINK format Genome Wide Association Studies -
Face-to-faceWorkshops and coursesImplementation of Data Management Plans & Data Stewardship in practice
11 September 2018 @ 17:30 - 19:00
Athens, Greece
Face-to-face
Data submission, annotation, and curation Data management data management data stewardship eLearning EeLP -
Face-to-faceWorkshops and coursesELIXIR Data Carpentry - Ljubljana
29 - 30 January 2019
Ljubljana, Slovenia
Face-to-face
Computer science Data management Genomics EeLP eLearning genomics Data carpentry -
Face-to-faceWorkshops and coursesELIXIR Software Carpentry - Ljubljana
16 - 17 May 2019
Ljubljana, Slovenia
Face-to-face
Computer science Data management Statistics and probability EeLP eLearning Statistics R-programming Software Carpentry -
OnlineWorkshops and coursesHow to share text mining results in biology
9 July 2019
Online
Text mining Literature and language Literature search Scientific communication Scientific presentation Data presentation Research presentation Protein Data Bank in Europe Ensembl UniProt: The Universal Protein Resource AlphaFold Database Proteins (proteins) DNA & RNA (dna-rna) Variant prediction Fermentation Microbial ecosystems webinar Antimicrobial resistance BLAST Open Targets Platform Cross domain (cross-domain) Chemical biology (chemical-biology) Drug discovery Drug target identification UniRule ARBA Automated annotation MetaboLights: Metabolomics repository and reference database Chemical Entities of Biological Interest ChEBI Metabolites Molecular building blocks of life Human Cell Atlas Data Coordination Platform Single-cell transcriptomics HCA data portal Programmatic access API Python Complex Portal macromolecular assembly InterPro Boolean modelling Europe PubMed Central Literature (literature) Open access Protein Data Bank in Europe - Knowledge Base 3D structure DeepMind Artificial intelligence AI Structure prediction cancer Boolean Ensembl Genomes European Nucleotide Archive Data archive Raw sequencing data RNAcentral Non-coding RNA ncRNA GPU Data protection Job dispatcher Bioimage analysis resource Accessibility Missense variation Biostatistics Rfam non-coding RNA Infernal software Sequence annotation Root microbiome Abiotic stress land management Plant genotype Plant webinar series HPC database development cross-linked databases Plant database data infrastructure Plant breeding Data standards data managemnet data sharing Hyb-Seq method Flowering plants Crop improvement Pangenomics Pangenomes Virtual humans Drug-target identification plant-microbe interactions Spatial transcriptomics Plant research Drug targets Machine learning Mathematical modelling plant science Data integration plant-environment interaction Phenotyping field phenotyping Deep phenotyping EOSC-Life NHGRI-EBI GWAS Catalog clinical data genome-wide association plants European Variation Archive EVA Variant clusters Variant data annotation Constraint-based metabolic modelling UniProt knowledgebase protein variant impact disease-associated protein variants Bioethics FAIR principles ELSI cohort data translational research BioModels database Mathematical modeling Reproducibity Systems biology models workflows federated analysis polygenic risk scores IntAct Molecular Interaction Database PSICQUIC IMEx Complex portal Agent-based modelling Macrophages Tumorigenesis Training (Training) On-demand teaching introduction Building blocks Data analysis COSMIC Cancer mutation Somatic mutation ChEMBL: Bioactive data for drug discovery Chemical compounds drug-like molecules Chemogenomics Biocurator Programming Data management Green Algorithms Open data Environmental impact Carbon footprint HPC workflows Orchestrator Gene expression (gene-expression) Chemosensitivity assay Experimental protocols Drug screening MICHA European Genome-phenome Archive EGA restricted access UniProt Introduction UniProtKB Proteome genes Introductory GDPR Data security Expression Atlas Pfam protein sequence search Domain architecture Protein taxonomy AI structure prediction COVID-19 Coronavirus COVID-19 Data Portal Virology Computational simulations Signalling prior knowledge Cancer therapy response PRIDE: The Proteomics Identifications Database protein structure prediction AI system Competency framework Publication Journal club Preprints Aggregated view Structural similarity PPI networks Cell signaling Malaria Biological networks Graph theory Building biological networks Biocuration Information Manager Industry Rare-variant Variant association SKAT Sensitive data Human cohort data FAIR Gene Ontology International Mouse Phenotyping Consortium Portal Biocurators biocuration programming skills training career development Cell-level simulations Personalised medicine PerMedCoE Single cell Phenotype Genome-Wide Association Studies Genome Signaling networks BioImage Archive REST Funding UniChem: Chemical Structure Cross-referencing Bioactivity data Drug-like compounds BioStudies Database EBI Search Finding data World Sight Day Eye development Eye disease Open software Metadata Web Services Dbfetch Database fetch programmatic access Open Targets Protein structure prediction Colab notebooks RESTful API RESTful Web Services PDBe Ligand binding sites Protein families Protein functional analysis Protein sequences Data curation Biomolecular sequences Cytoscape PsyArXiv Psychology Bioactive data Drug-like molecules Biological studies Literature Neurodegenerative disorders Data sharing Molecular visualisation Molecular structures Electron density maps EBI BioSamples Database BioSamples Data Use Ontology DUO ELIXIR Ensembl Variant Effect Predictor Variant Effect Predictor VEP Variation data Annotation SciLite annotation MGnify Microbiome Environment Proteomes Variant data Publications Citations UniProt Disease Portal ENA Sequence Search Reactome pathways database ReactomeGSA Pathway analysis Single cell RNA-seq Multiomics Data visualisation GitHub Graph database protein Gene function Mouse models IMPC Mammalian Phenotype Ontology Mouse knockouts Xenograft models Cancer EurOPDX InterProScan PDBe-KB Interpro API Europe PMC PubMed Protein function Preprints SARS-CoV-2 Genome browser Vertebrate genomes Comparative genomics Drug-like properties eQTL Catalogue Single Cell Expression Atlas SCEA Life cell by cell Introduction to single cell expression atlas PDX UniParc UniRef Genes GENCODE Gene annotation Clinical genomics Genome3D Genome annotations Protein domain prediction -
OnlineWorkshops and coursesBioStudies database: aggregating all outputs of a life sciences study
17 July 2019
Online
Data retrieval Literature and language Experimental design and studies Scientific communication Scientific presentation Data presentation Research presentation Protein Data Bank in Europe Ensembl UniProt: The Universal Protein Resource AlphaFold Database Proteins (proteins) DNA & RNA (dna-rna) Variant prediction Fermentation Microbial ecosystems webinar Antimicrobial resistance BLAST Open Targets Platform Cross domain (cross-domain) Chemical biology (chemical-biology) Drug discovery Drug target identification UniRule ARBA Automated annotation MetaboLights: Metabolomics repository and reference database Chemical Entities of Biological Interest ChEBI Metabolites Molecular building blocks of life Human Cell Atlas Data Coordination Platform Single-cell transcriptomics HCA data portal Programmatic access API Python Complex Portal macromolecular assembly InterPro Boolean modelling Europe PubMed Central Literature (literature) Open access Protein Data Bank in Europe - Knowledge Base 3D structure DeepMind Artificial intelligence AI Structure prediction cancer Boolean Ensembl Genomes European Nucleotide Archive Data archive Raw sequencing data RNAcentral Non-coding RNA ncRNA GPU Data protection Job dispatcher Bioimage analysis resource Accessibility Missense variation Biostatistics Rfam non-coding RNA Infernal software Sequence annotation Root microbiome Abiotic stress land management Plant genotype Plant webinar series HPC database development cross-linked databases Plant database data infrastructure Plant breeding Data standards data managemnet data sharing Hyb-Seq method Flowering plants Crop improvement Pangenomics Pangenomes Virtual humans Drug-target identification plant-microbe interactions Spatial transcriptomics Plant research Drug targets Machine learning Mathematical modelling plant science Data integration plant-environment interaction Phenotyping field phenotyping Deep phenotyping EOSC-Life NHGRI-EBI GWAS Catalog clinical data genome-wide association plants European Variation Archive EVA Variant clusters Variant data annotation Constraint-based metabolic modelling UniProt knowledgebase protein variant impact disease-associated protein variants Bioethics FAIR principles ELSI cohort data translational research BioModels database Mathematical modeling Reproducibity Systems biology models workflows federated analysis polygenic risk scores IntAct Molecular Interaction Database PSICQUIC IMEx Complex portal Agent-based modelling Macrophages Tumorigenesis Training (Training) On-demand teaching introduction Building blocks Data analysis COSMIC Cancer mutation Somatic mutation ChEMBL: Bioactive data for drug discovery Chemical compounds drug-like molecules Chemogenomics Biocurator Programming Data management Green Algorithms Open data Environmental impact Carbon footprint HPC workflows Orchestrator Gene expression (gene-expression) Chemosensitivity assay Experimental protocols Drug screening MICHA European Genome-phenome Archive EGA restricted access UniProt Introduction UniProtKB Proteome genes Introductory GDPR Data security Expression Atlas Pfam protein sequence search Domain architecture Protein taxonomy AI structure prediction COVID-19 Coronavirus COVID-19 Data Portal Virology Computational simulations Signalling prior knowledge Cancer therapy response PRIDE: The Proteomics Identifications Database protein structure prediction AI system Competency framework Publication Journal club Preprints Aggregated view Structural similarity PPI networks Cell signaling Malaria Biological networks Graph theory Building biological networks Biocuration Information Manager Industry Rare-variant Variant association SKAT Sensitive data Human cohort data FAIR Gene Ontology International Mouse Phenotyping Consortium Portal Biocurators biocuration programming skills training career development Cell-level simulations Personalised medicine PerMedCoE Single cell Phenotype Genome-Wide Association Studies Genome Signaling networks BioImage Archive REST Funding UniChem: Chemical Structure Cross-referencing Bioactivity data Drug-like compounds BioStudies Database EBI Search Finding data World Sight Day Eye development Eye disease Open software Metadata Web Services Dbfetch Database fetch programmatic access Open Targets Protein structure prediction Colab notebooks RESTful API RESTful Web Services PDBe Ligand binding sites Protein families Protein functional analysis Protein sequences Data curation Biomolecular sequences Cytoscape PsyArXiv Psychology Bioactive data Drug-like molecules Biological studies Literature Neurodegenerative disorders Data sharing Molecular visualisation Molecular structures Electron density maps EBI BioSamples Database BioSamples Data Use Ontology DUO ELIXIR Ensembl Variant Effect Predictor Variant Effect Predictor VEP Variation data Annotation SciLite annotation Text mining MGnify Microbiome Environment Proteomes Variant data Publications Citations UniProt Disease Portal ENA Sequence Search Reactome pathways database ReactomeGSA Pathway analysis Single cell RNA-seq Multiomics Data visualisation GitHub Graph database protein Gene function Mouse models IMPC Mammalian Phenotype Ontology Mouse knockouts Xenograft models Cancer EurOPDX InterProScan PDBe-KB Interpro API Europe PMC PubMed Protein function Preprints SARS-CoV-2 Genome browser Vertebrate genomes Comparative genomics Drug-like properties eQTL Catalogue Single Cell Expression Atlas SCEA Life cell by cell Introduction to single cell expression atlas PDX UniParc UniRef Genes GENCODE Gene annotation Clinical genomics Genome3D Genome annotations Protein domain prediction
Note, this map only displays events that have geolocation information in
TeSS.
For the complete list of events in TeSS, click the grid tab.